Juan, thanks for your input. My apologies for not giving details earlier as I suspected it was a local machine issue rather than one with the excellent PI software.
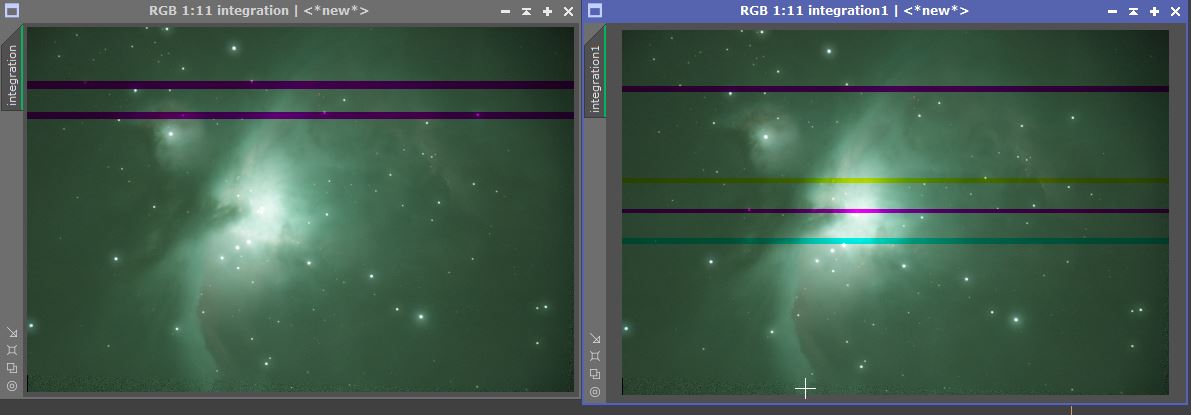
Some history. I first noticed this effect back in March when integrating big datasets from my QHY163C camera, acquired using Sequence Generator Lite. The 107 fit files were then calibrated with bias, dark and flats and saved in xisf format, cosmetic corrected, debayered and star aligned. I then created local normalisation files with the default settings. Integration settings: with Combination: Average, Normalisation: Local normalisation, Weights: Noise evaluation with pixel rejection: Percentile clipping, with local normalisation. Other setting at default. Screenshot on OneDrive. On integration, I found the result had these lines of a dropped colour. I repeated the integration 5 times (I think) before I got one without these artefacts. At that point I stopped and moved on.
Moving forward, last week I took 6 images on a Canon 750D camera in CR2 format. Manually triggered so no control software. On integration, a similar thing happened - banding as first image I posted. After 4 repeat integrations, I had a successful one and stopped. My suspicion that the effect was due to the large number [a rarity for me] of files was no longer possible, so I enquired about this problem on the forum.
Given the comments, I wondered if this was a swap file issue so I decided to try and move the swap files to a different drive but saw the warning pop-up about not to do this on a mechanical drive. Instead, I reduced the number of swap files [C:\TEMP ] on the SSD to 8 - from the 12 which had seemed [marginally] better from the PI Benchmark process. [Screenshot of Preferences tab on OneDrive] Yesterday I tried to reproduce this banding with the 6 file set and I find I cannot after 13 attempts. I have now integrated the 107 fileset over 10 times with the 8 swap files and again no banding issues. Switching back to 12, second and third reintegration gave bands. Sometimes the bands are not obvious within the overall green bias [did I miss them on the 8 swapfile tests?] and only become clear with a subtractive ABE [default settings]. Closer inspection reveals that star colours are incorrect on original integration. This morning I found some integrations with 8 swapfiles giving banding.
I now have a 20 file subset which show fairly repeatedly, but not every time, the banding issue on integration with 12 swap files and occasionally 8 swap files.. The problem seemed much more frequent on the 12 swap file setting.
I hope this is what you need. I have uploaded the 20 subs [zipped] to onedrive together with a screenshot of the integration settings and four example integration results from the 20 file subset.
The onedrive address is
https://1drv.ms/u/s!Ajb0X0vgvXvUgdocPzxT9x5syxt-hA?e=tRsdwx
System:
Processor: Intel Core i7-8700@ 3.2GHz,
Mobo: B360M AORUS Gaming 3 with 32GB ram,
Graphics Card: NVidea GeForce GTX 1050 Ti
System Disk: Samsung SSD 860 EVO M.2 1TB
OS: Win10 Home 64bit build 19041.572
PI version: 1.8.8-6